Research Paper:
Brightness-Sensitive Generative Adversarial Network Using a Chained Extension Framework for PET-to-CT Medical Image Synthesis
Xiaoyu Deng*,†
 , Kouki Nagamune**
, Kouki Nagamune**
 , and Hiroki Takada*
, and Hiroki Takada*
*Graduate School of Engineering, University of Fukui
3-9-1 Bunkyo, Fukui, Fukui 910-0017, Japan
†Corresponding author
**Department of Electronics and Computer Science, Graduate School of Engineering, University of Hyogo
2167 Shosha, Himeji, Hyogo 671-2280, Japan
Multimodal medical imaging is pivotal for early disease screening; however, its deployment is often constrained by limited resources. While deep learning-based synthesis cannot replace clinical imaging, high-fidelity cross-modal translation can provide actionable prior information for preliminary assessment. To this end, we introduce a chained extension framework that scales model capacity and precision by linking multiple encoder–decoder modules. Starting from a minimal encoder–decoder backbone, we construct a triple-stage generative adversarial network and integrate a brightness-sensitive loss that reweights luminance-dependent errors. This staged design decomposes positron emission tomography to computed tomography translation into complementary subtasks targeting structural consistency, texture enhancement, and key-region refinement. Comprehensive experiments indicate that the proposed approach generates synthetic CT images that closely match reference CT scans, visually and quantitatively, achieving a structural similarity index of 0.85, peak signal-to-noise ratio of 23.31 dB, and mean absolute error of 6.93×10-2. Thus, our framework is feasible as an assistive tool for early screening workflows in resource-limited settings. Moreover, the staged training strategy, coupled with brightness-aware weighting, mitigates common optimization plateaus in cross-modal synthesis, suggesting a principled path toward further gains in fidelity and robustness.
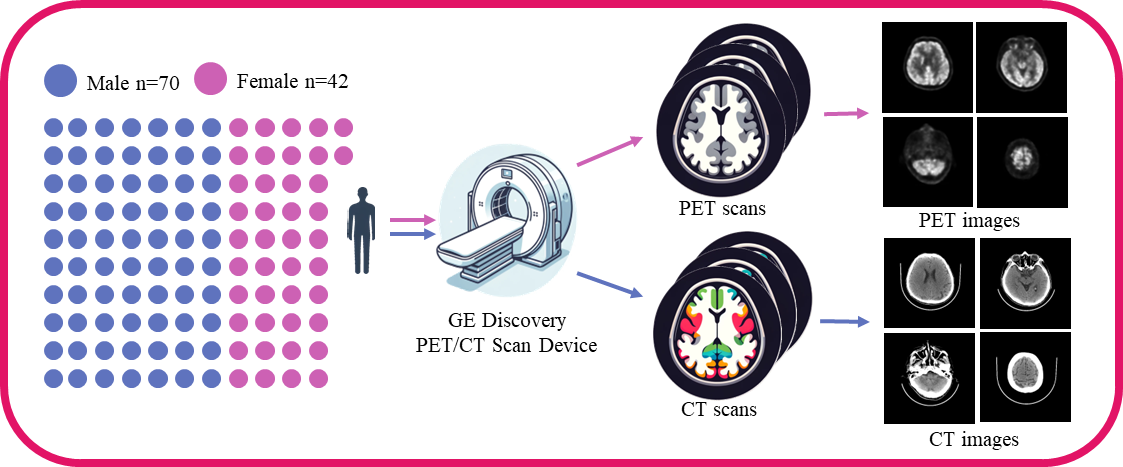
Data acquisition and preprocessing steps
- [1] H. Schöder, Y. E. Erdi, S. M. Larson, and H. W. D. Yeung, “PET/CT: A new imaging technology in nuclear medicine,” European J. of Nuclear Medicine and Molecular Imaging, Vol.30, No.10, pp. 1419-1437, 2003. https://doi.org/10.1007/s00259-003-1299-6
- [2] M. W. Saif, I. Tzannou, N. Makrilia, and K. Syrigos, “Role and Cost Effectiveness of PET/CT in Management of Patients with Cancer,” The Yale J. of Biology and Medicine, Vol.83, No.2, pp. 53-65, 2010.
- [3] O. Ronneberger, P. Fischer, and T. Brox, “U-Net: Convolutional Networks for Biomedical Image Segmentation,” N. Navab, J. Hornegger, W. M. Wells, and A. F. Frangi (Eds.), Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015, Lecture Notes in Computer Science, Vol.9351, pp. 234-241, 2015. https://doi.org/10.1007/978-3-319-24574-4_28
- [4] P. Isola, J.-Y. Zhu, T. Zhou, and A. A. Efros, “Image-to-Image Translation with Conditional Adversarial Networks,” 2017 IEEE Conf. on Computer Vision and Pattern Recognition (CVPR), pp. 5967-5976, 2017. https://doi.org/10.1109/CVPR.2017.632
- [5] T. Wang, Y. Lei, Y. Fu, J. F. Wynne, W. J. Curran, T. Liu, and X. Yang, “A review on medical imaging synthesis using deep learning and its clinical applications,” J. of Applied Clinical Medical Physics, Vol.22, Issue 1, pp. 11-36, 2021. https://doi.org/10.1002/acm2.13121
- [6] S. Kaji and S. Kida, “Overview of image-to-image translation by use of deep neural networks: Denoising, super-resolution, modality conversion, and reconstruction in medical imaging,” Radiological Physics and Technology, Vol.12, No.3, pp. 235-248, 2019. https://doi.org/10.1007/s12194-019-00520-y
- [7] M. E. Rayed, S. M. S. Islam, S. I. Niha, J. R. Jim, M. M. Kabir, and M. Mridha, “Deep learning for medical image segmentation: State-of-the-art advancements and challenges,” Informatics in Medicine Unlocked, Vol.47, Article No.101504, 2024. https://doi.org/10.1016/j.imu.2024.101504
- [8] K. Suzuki, “Overview of deep learning in medical imaging,” Radiological Physics and Technology, Vol.10, No.3, pp. 257-273, 2017. https://doi.org/10.1007/s12194-017-0406-5
- [9] V. Sevetlidis, M. V. Giuffrida, and S. A. Tsaftaris, “Whole Image Synthesis Using a Deep encoder–decoder Network,” S. A. Tsaftaris, A. Gooya, A. F. Frangi, and J. L. Prince (Eds.), “Simulation and Synthesis in Medical Imaging,” Lecture Notes in Computer Science, Vol.9968, pp. 127-137, 2016. https://doi.org/10.1007/978-3-319-46630-9_13
- [10] F. Hashimoto, M. Ito, K. Ote, T. Isobe, H. Okada, and Y. Ouchi, “Deep learning-based attenuation correction for brain PET with various radiotracers,” Annals of Nuclear Medicine, Vol.35, No.6, pp. 691-701, 2021. https://doi.org/10.1007/s12149-021-01611-w
- [11] J. Zhang, Z. Cui, C. Jiang, J. Zhang, F. Gao, and D. Shen, “Mapping in Cycles: Dual-Domain PET-CT Synthesis Framework with Cycle-Consistent Constraints,” L. Wang, Q. Dou, P. T. Fletcher, S. Speidel, and S. Li (Eds.), “Medical Image Computing and Computer Assisted Intervention (MICCAI 2022),” Lecture Notes in Computer Science, Vol.13436, pp. 758-767, 2022. https://doi.org/10.1007/978-3-031-16446-0_72
- [12] Y. Lecun, L. Bottou, Y. Bengio, and P. Haffner, “Gradient-based learning applied to document recognition,” Proc. of the IEEE, Vol.86, Issue 11, pp. 2278-2324, 1998. https://doi.org/10.1109/5.726791
- [13] J. Pons, S. Pascual, G. Cengarle, and J. Serrà, “Upsampling Artifacts in Neural Audio Synthesis,” 2021 IEEE Int. Conf. on Acoustics, Speech and Signal Processing (ICASSP 2011), pp. 3005-3009, 2021. https://doi.org/10.1109/ICASSP39728.2021.9414913
- [14] I. J. Goodfellow, J. Pouget-Abadie, M. Mirza, B. Xu, D. Warde-Farley, S. Ozair, A. Courville, and Y. Bengio, “Generative Adversarial Nets,” Advances in Neural Information Processing Systems, Vol.27, 2014.
- [15] C. Guo, M. Szemenyei, Y. Yi, W. Wang, B. Chen, and C. Fan, “SA-UNet: Spatial Attention U-Net for Retinal Vessel Segmentation,” 2020 25th Int. Conf. on Pattern Recognition (ICPR), pp. 1236-1242, 2021. https://doi.org/10.1109/ICPR48806.2021.9413346
- [16] O. Petit, N. Thome, C. Rambour, L. Themyr, T. Collins, and L. Soler, “U-Net Transformer: Self and Cross Attention for Medical Image Segmentation,” C. Lian, X. Cao, I. Rekik, X. Xu, and P. Yan (Eds.), “Machine Learning in Medical Imaging,” Lecture Notes in Computer Science, Vol.12966, pp. 267-276, 2021. https://doi.org/10.1007/978-3-030-87589-3_28
- [17] A. Singh, J. Kwiecinski, S. Cadet, A. Killekar, E. Tzolos, M. C. Williams, M. R. Dweck, D. E. Newby, D. Dey, and P. J. Slomka, “Automated nonlinear registration of coronary PET to CT angiography using pseudo-CT generated from PET with generative adversarial networks,” J. of Nuclear Cardiology, Vol.30, Issue 2, pp. 604-615, 2023. https://doi.org/10.1007/s12350-022-03010-8
- [18] X. Dong, T. Wang, Y. Lei, K. Higgins, T. Liu, W. J. Curran, H. Mao, J. A. Nye, and X. Yang, “Synthetic CT generation from non-attenuation corrected PET images for whole-body PET imaging,” Physics in Medicine & Biology, Vol.64, No.21, Article No.215016, 2019. https://doi.org/10.1088/1361-6560/ab4eb7
- [19] J. Li, Y. Wang, Y. Yang, X. Zhang, Z. Qu, and S. Hu, “Small animal PET to CT image synthesis based on conditional generation network,” 2021 14th Int. Congress on Image and Signal Processing, BioMedical Engineering and Informatics (CISP-BMEI), 2021. https://doi.org/10.1109/CISP-BMEI53629.2021.9624232
- [20] H. Wang, X. Wang, F. Liu, G. Zhang, G. Zhang, Q. Zhang, and M. L. Lang, “DSG-GAN: A dual-stage-generator-based GAN for cross-modality synthesis from PET to CT,” Computers in Biology and Medicine, Vol.172, Article No.108296, 2024. https://doi.org/10.1016/j.compbiomed.2024.108296
- [21] J. Li, Z. Qu, Y. Yang, F. Zhang, M. Li, and S. Hu, “TCGAN: A transformer-enhanced GAN for PET synthetic CT,” Biomedical Optics Express, Vol.13, Issue 11, pp. 6003-6018, 2022. https://doi.org/10.1364/BOE.467683
- [22] X. Chen, S. Luo, C.-M. Pun, and S. Wang, “MedPrompt: Cross-modal Prompting for Multi-task Medical Image Translation,” Z. Lin, M.-M. Cheng, R. He, K. Ubul, W. Silamu, H. Zha, J. Zhou, and C.-L. Liu (Eds.), “Pattern Recognition and Computer Vision,” Lecture Notes in Computer Science, Vol.15044, pp. 61-75, 2025. https://doi.org/10.1007/978-981-97-8496-7_5
- [23] J. Liang, H. Zeng, and L. Zhang, “High-Resolution Photorealistic Image Translation in Real-Time: A Laplacian Pyramid Translation Network,” Proc. of 2021 IEEE/CVF Conf. on Computer Vision and Pattern Recognition (CVPR), pp. 9387-9395, 2021. https://doi.org/10.1109/CVPR46437.2021.00927
- [24] O. Dalmaz, M. Yurt, and T. Cukur, “ResViT: Residual Vision Transformers for Multimodal Medical Image Synthesis,” IEEE Trans. on Medical Imaging, Vol.41, Issue 10, pp. 2598-2614, 2022. https://doi.org/10.1109/TMI.2022.3167808
- [25] F. Gao, T. Wu, X. Chu, H. Yoon, Y. Xu, and B. Patel, “Deep Residual Inception Encoder–Decoder Network for Medical Imaging Synthesis,” IEEE J. of Biomedical and Health Informatics, Vol.24, Issue 1, pp. 39-49, 2020. https://doi.org/10.1109/JBHI.2019.2912659
- [26] M.-Y. Liu, T. Breuel, and J. Kautz, “Unsupervised Image-to-Image Translation Networks,” Proc. of the 31st Int. Conf. on Neural Information Processing Systems (NIPS’17), pp. 700-708, 2017.
- [27] X. Huang, M.-Y. Liu, S. Belongie, and J. Kautz, “Multimodal Unsupervised Image-to-Image Translation,” V. Ferrari, M. Hebert, C. Sminchisescu, and Y. Weiss (Eds.), “Proc. of the European Conf. on Computer Vision (ECCV2018),” Lecture Notes in Computer Science, Vol.11207, pp. 179-196, 2018. https://doi.org/10.1007/978-3-030-01219-9_11
- [28] M.-Y. Liu, X. Huang, A. Mallya, T. Karras, T. Aila, J. Lehtinen, and J. Kautz, “Few-Shot Unsupervised Image-to-Image Translation,” Proc. of 2019 IEEE/CVF Int. Conf. on Computer Vision (ICCV), pp. 10550-10559, 2019. https://doi.org/10.1109/ICCV.2019.01065
- [29] M. K. Sherwani and S. Gopalakrishnan, “A systematic literature review: Deep learning techniques for synthetic medical image generation and their applications in radiotherapy,” Frontiers in Radiology, Vol.4, Article No.1385742, 2024. https://doi.org/10.3389/fradi.2024.1385742
 This article is published under a Creative Commons Attribution-NoDerivatives 4.0 Internationa License.
This article is published under a Creative Commons Attribution-NoDerivatives 4.0 Internationa License.